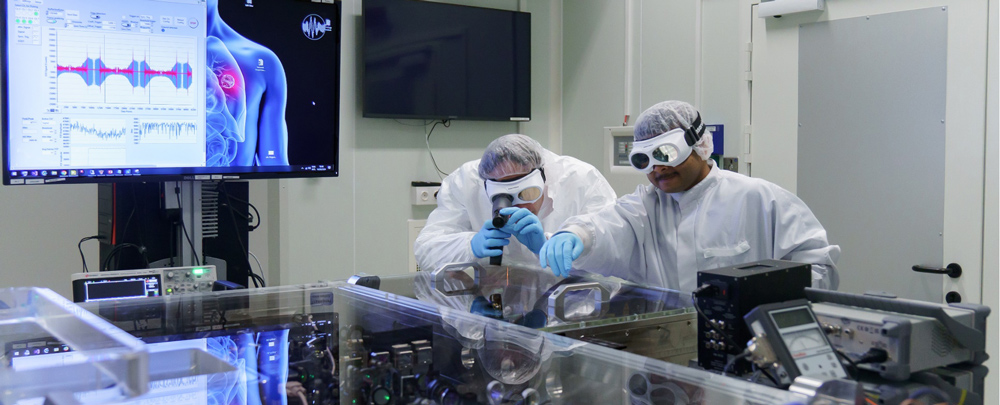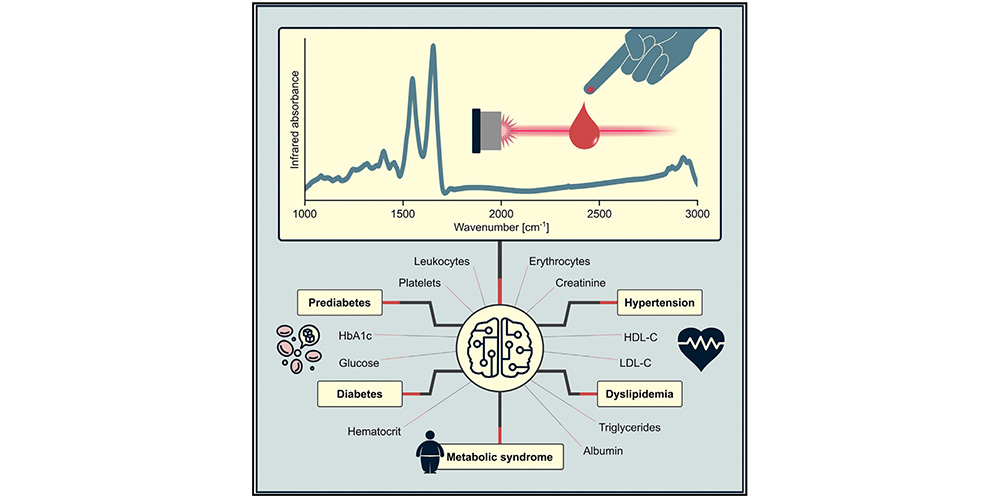
Our research combines advanced InfraRed spectroscopy technologies with multi-omics, clinical diagnostics, and computational data analyses to develop novel molecular fingerprinting methods for human health assessment and early disease detection. The research builds on several clinical studies for which thousands of individuals contribute blood samples, resulting in rapidly growing datasets with rich molecular measurements and detailed clinical characterization. We are an interdisciplinary team with expertise spanning physics, data science, molecular science, and clinical research. Our main question is to understand how subtle molecular changes in blood relate to health states, disease onset, and longitudinal health trajectories, with applications in common diseases and cancer. To integrate our findings and generate new predictions and hypotheses, we use machine learning and systems analysis frameworks. Further information is available at www.attoworld.de/bird
Research area: Projects will focus on analysis of large-scale molecular datasets applying machine learning and data analysis. Research directions include the development of robust approaches for data standardization, improving model generalization across heterogeneous datasets, and extracting clinically meaningful insights from complex molecular data.
- a strong academic background in informatics, bioinformatics, data engineering and analytics, statistics, computational physics, or a related discipline;
- solid expertise in statistics, machine learning, or data analytics;
- programming experience, ideally in Python;
- strong analytical and problem-solving abilities;
- a genuine interest in interdisciplinary biomedical research.
Prior experience in molecular biology, biophysics, or chemistry is advantageous but not essential. More important is the motivation to work across disciplines and to engage with biologically and clinically relevant questions. We value diversity and strongly encourage applications from all qualified individuals.
- a friendly, vibrant, and collaborative environment with regular seminars, journal clubs, core lab funding;
- extensive collaborative opportunities with other lab members, labs, and across institutes;
- support in developing leadership, computational, and experimental skills;
- structured mentoring, with opportunities to co-supervise, lead projects, and help shape the group’s scientific direction.
Applications, including a letter of motivation, a CV and the names and contact details of two referees should be sent directly to Mihaela Zigman and Tarek Eissa (mihaela.zigman@mpq.mpg.de; tarek.eissa@mpq.mpg.de).
The group is based at Garching Forschungszentrum. For PhD applicants, formal supervision can be arranged through LMU Physics or TUM School of Computation Information and Technology (CIT).
- Eissa et al., “Plasma infrared fingerprinting with machine learning enables single-measurement multi-phenotype health screening.” Cell Reports Medicine 5, 101625 (2024).
- Eissa et al., “CODI: enhancing machine learning-based molecular profiling through contextual out-of-distribution integration.” PNAS Nexus 3, pgae449 (2024).
- Eissa et al., “The perils of molecular interpretations from vibrational spectra of complex samples.” Angewandte Chemie Int. Edition 63, e202411596 (2024).
- Kepesidis et al., “Electric-field molecular fingerprinting to probe cancer.” ACS Central Science 11 (4), 560-573 (2025).
Download: Job Description (PDF)
less